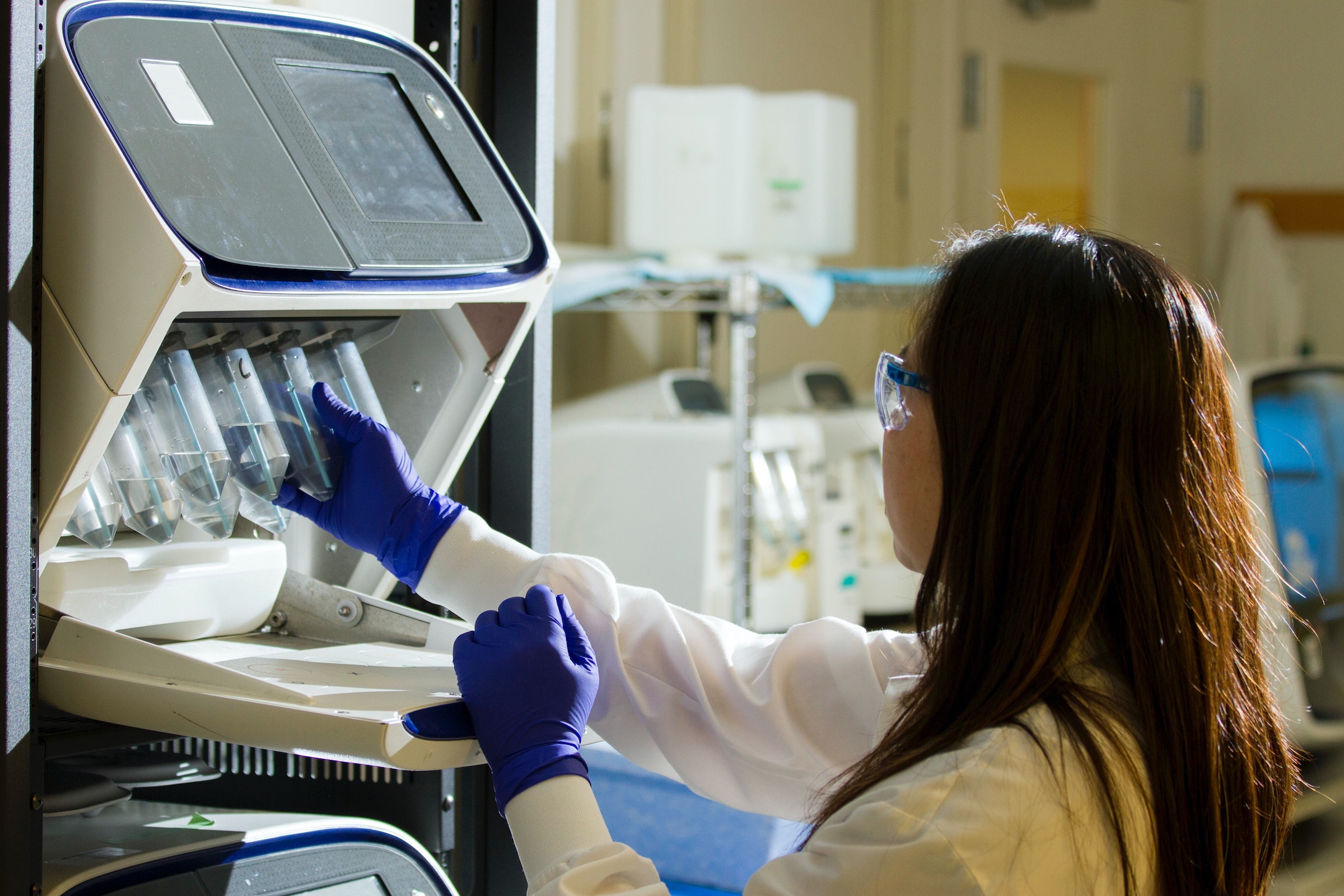
COVID-19 has compelled the global research community to push the boundaries of genetic sequencing in virus identification and tracing.
For all the problems that the pandemic presents, we have never been in a better position to explore the applications of genomic epidemiology. Advances in technology have allowed us to sequence, track, and map genes faster and more thoroughly than ever before — and then immediately share our findings with researchers halfway across the world.
However, our advancements have not given us a “golden ticket” to understanding COVID-19. Instead, COVID-19 has compelled us to explore how our progress in genomic epidemiology can be strategically applied in the resolution of a global health crisis. In this article, I will address one such application — virus research — and consider how our COVID-19 explorations could inform future approaches to public health crises.
Over the past several months, researchers have used genomic epidemiology research to better understand COVID-19, its spread, and how both should inform local containment measures.
At this point, it may be worth providing some context on the research process. The practice of analyzing viral genomes to trace their spread has been around for decades. The core idea is simple. During the replication process, a virus’s protein machinery will make a mistake and thereby create a mutated strain. By counting these mutations, genomic epidemiologists can track the virus’s travels and evolution.
When researchers assess the rate of mutation, the number of mutations found in local virus strains, and sequences seen in the believed country of origin, they can make an informed estimate of how long the virus has been present in a community and how many people in the region are likely infected with the virus. This information can then be passed on to public health authorities and used to inform public health response.
For a practical example, we can turn to Washington State. In March, the open-source genome and pathogen research project Nextstrain compared two locally-sampled SARS-CoV-2 genomes that were collected six weeks apart. After realizing that both genomes featured a mutation rarely seen in genomes from Wuhan — the believed source of the Washington outbreak — Nextstrain scientists concluded that local transmission of the virus had to be occurring in the Pacific Northwest.
Nextstrain shared this data with public health authorities. Within a few days, both the governor and the mayor of Seattle had declared a state of emergency and indicated localized spread as proof of the elevated risk to public health.
Nextstrain’s research demonstrates how genomic epidemiology investigations can inform large-scale responses to a pandemic. A similar tactic can also be applied to managing micro-outbreaks. In the UK, a team of clinicians and researchers affiliated with the University of Cambridge and the Cambridge University Hospitals NHS Foundation Trust (CUH) analyzed the virus’s genetic code to see if cases found in the hospital shared a common source. This analysis, in turn, allowed the hospital’s staff to investigate outbreaks within their facilities and limit the risk of further infection.
As writers for Medical Xpress reported on the initiative, “the introduction of a screening programme that involved repeat testing of staff, has helped the hospital to investigate clusters of COVID-19 infections, informing infection control measures and breaking chains of transmission. This has helped reduce the number of hospital-acquired infections, ensuring maximum safety for patients and staff as the NHS aims to re-start other services.”
“The existence of clusters of infection in specific areas of the hospital shows the potential for staff and patients to become infected within the hospital environment,” Dr. Steve Baker, a professor at the Cambridge Institute of Therapeutic Immunology & Infectious Disease, told reporters. “If left unchecked, these clusters could lead to self-sustaining outbreaks. Frequent testing at CUH allowed us to spot these clusters quickly and stop any further transmission.”
The importance of finding and addressing COVID-19 outbreaks before they contribute to significant disease spread cannot be understated. If CUH’s approach — or, to think more broadly, genomic tracing methods as a whole — could be replicated at hospitals and in public health offices within virus hotspots, it could prove invaluable to tamping down infection rates and resolving the crisis.